Notes on Biophysical Chemistry (MDC)
We provide notes on all MDC Papers for Chemistry such as Biochemistry, Foods and nutrition, Environmental Chemistry.
Unit I: Fundamentals of Biological Macromolecules
Biomacromolecules are large
biological molecules that are essential for life. They are typically composed
of smaller molecular subunits and play crucial roles in the structure and
function of cells.
The four major types of
biomacromolecules are:
1.Carbohydrates – Made up of
monosaccharides (simple sugars), these molecules provide energy and structural
support. Example: Starch, glycogen, and cellulose.
2.Proteins – Composed of amino acids,
proteins serve various functions including enzymatic activity, structural support,
and signaling. Example: Hemoglobin, enzymes, and collagen.
3.Nucleic Acids – Made up of
nucleotides, these molecules store and transmit genetic information. Example:
DNA and RNA.
4.Lipids
– Although not always considered macromolecules due to their non-polymeric
nature, lipids are crucial for cell membranes, energy storage, and signaling.
Example: Fats, phospholipids, and steroids.
Click here for previous year question papers
Click here for MDC Biochemistry
Proteins: Definition & Structure
Proteins are biological
macromolecules made up of long chains of amino acids linked by peptide bonds.
They are essential for nearly all biological processes, playing structural,
enzymatic, and regulatory roles in cells.
Chemical Structure of Amino Acids
Amino acids are the building blocks
of proteins. They have a general structure consisting of a central α-carbon
bonded to four groups:
1.Amino group (-NH₂) → Basic in nature
2.Carboxyl group (-COOH) → Acidic in
nature
3.Hydrogen atom (H)
4.R-group (Side Chain) → Unique for
each amino acid
General Structure:
H2N−CH(R)−COOH
The R-group determines the chemical
properties and classification of the amino acid.
Classification of Amino Acids Based
on R-Groups
1.Nonpolar (Hydrophobic) Amino Acids
Contain alkyl or aromatic R-groups
Examples: Glycine, Alanine, Valine,
Leucine, Isoleucine, Phenylalanine
2.Polar (Uncharged) Amino Acids
Contain hydroxyl (-OH), amide
(-CONH₂), or thiol (-SH) groups
Examples: Serine, Threonine,
Tyrosine, Asparagine, Glutamine
3.Acidic (Negatively Charged) Amino
Acids
Contain extra carboxyl (-COOH) group
Examples: Aspartic acid, Glutamic
acid
4.Basic (Positively Charged) Amino
Acids
Contain extra amino (-NH₂) group
Examples: Lysine, Arginine, Histidine
Zwitterion Form
Amino acids exist as zwitterions, meaning they have both a positive (-NH₃⁺) and negative (-COO⁻) charge.
Example: H3N+−CH(R)−COO−
This property affects solubility and reactivity.
Ionization States of Amino Acids
The ionization state of an amino acid depends on the pH of the solution and its pKa values.
a) At Low pH (Acidic Environment)
- The carboxyl group remains protonated (-COOH).
- The amino group is protonated (-NH₃⁺).
- The overall charge is positive.
b) At Neutral pH
(Zwitterion Form, pH ≈ 7)
- The carboxyl group loses a proton
(-COO⁻).
- The amino group remains
protonated (-NH₃⁺).
- The molecule has no
net charge (Zwitterion: a species with both positive
and negative charges).
c) At High pH
(Basic Environment)
- The carboxyl group remains
deprotonated (-COO⁻).
- The amino group loses a proton
(-NH₂).
- The overall charge is negative.
3. Isoelectric Point (pI)
The isoelectric point (pI) is the pH at which the amino acid has no net
charge (exists primarily in the zwitterionic form). It is calculated as:
- For non-polar and uncharged polar amino acids:
- For acidic amino acids (Aspartic acid, Glutamic acid):
- For basic amino acids (Lysine, Arginine, Histidine):
Polypeptide
A polypeptide is a long, continuous
chain of amino acids linked together by peptide bonds. It serves as the
building block of proteins, which perform various biological functions in
cells.
Formation of Polypeptides
Peptide Bond Formation:
A peptide bond forms between the carboxyl
(-COOH) group of one amino acid and the amino (-NH₂) group of another.
This reaction is a condensation
reaction (removes a molecule of water, H₂O).
The resulting structure consists of
repeating amide (-CONH-) linkages.
Directionality:
N-Terminus (Amino End): The start of the polypeptide, containing a free amino (-NH₂) group.
C-Terminus (Carboxyl End): The end of
the polypeptide, containing a free carboxyl (-COOH) group.
Polypeptide: A simple chain of amino
acids, may not be functional.
Protein: A functional molecule that
may consist of one or more polypeptide chains folded into a specific 3D
structure.
Functions of Polypeptides
Enzymes (e.g., DNA polymerase)
Structural proteins (e.g., collagen,
keratin)
Hormones (e.g., insulin)
Transport proteins (e.g., hemoglobin)
Structure of Proteins
Proteins are made from 20 different amino acids, each with a unique side chain (R-group) that determines its properties. The structure of a protein can be described in four levels starting from the complex protein molecule to the unfolding of polypeptide chains and then to the detachment of different amino acids:
1.Quaternary Structure – The
arrangement of multiple polypeptide chains (subunits) in a functional protein
(e.g., hemoglobin has four subunits).
2.Tertiary Structure – The overall 3D shape of a single polypeptide, stabilized by interactions like hydrogen bonds, ionic bonds, disulfide bridges, and hydrophobic interactions.
3.Secondary Structure – Local folding patterns such as:
α-Helix (coiled structure)
β-Sheet (pleated sheet structure) These are stabilized by hydrogen bonds.
4.Primary Structure – The linear sequence of amino acids in a polypeptide chain.
Internal Rotation Angles:
In proteins, the internal
rotation angles (also known as torsion angles or dihedral angles)
refer to the angles between atoms that describe the conformation of the polypeptide
backbone and side chains. The most important torsion angles in
proteins are:
1. Phi (φ) angle
- Definition: Rotation
around the N–Cα bond (nitrogen to alpha-carbon).
- Affects how the polypeptide chain folds by rotating
the backbone.
2. Psi (ψ) angle
- Definition: Rotation
around the Cα–C' bond (alpha-carbon to carbonyl carbon).
- Along with φ, determines the secondary structure
(α-helix, β-sheet, etc.).
3. Omega (ω) angle
- Definition: Rotation
around the peptide bond (C'–N bond between amino acids).
- Usually fixed at 180° (trans) due to partial
double-bond character.
- Rarely, can be 0° (cis), mainly in proline.
4. Chi (χ) angles
- Definition: Torsion
angles around the side chain bonds starting from Cα.
- There can be multiple χ angles (χ1, χ2, etc.),
depending on the side chain.
- These determine the rotameric state of the side chain.
Click here to see the visual of rotation angles
Ramachandran Plot:
The Ramachandran Plot
is a graphical representation of the allowed conformations of amino acid
residues in protein structure based on the torsion angles φ (phi) and ψ
(psi).
Axes of the Plot
- X-axis: φ (phi)
angle (rotation around the N–Cα bond), ranges from –180° to +180°
- Y-axis: ψ (psi)
angle (rotation around the Cα–C' bond), also ranges from –180° to +180°
Purpose
- To identify sterically allowed regions of φ
and ψ angles.
- Helps predict secondary structures and
validate protein models.
Functions of Proteins
Proteins serve diverse roles in
biological systems:
1. As Structural Proteins – Provide support
(e.g., collagen in connective tissue, keratin in hair and nails).
2. As Enzymes – Speed up chemical reactions
(e.g., amylase, DNA polymerase).
3. As Transport Proteins – Carry molecules
(e.g., hemoglobin transports oxygen, membrane transporters).
4. Hormonal Proteins – Regulate
biological processes (e.g., insulin controls blood sugar).
5.Defensive Proteins – Protect against
diseases (e.g., antibodies in the immune system).
6.Contractile Proteins – Enable
movement (e.g., actin and myosin in muscles).
7.Storage Proteins – Store essential
substances (e.g., ferritin stores iron).
Sources of Proteins
Animal Sources: Meat, fish, eggs,
dairy.
Plant Sources: Beans, lentils, nuts,
soy.
Nucleic acids
Nucleic acids are biomolecules
essential for the storage, transmission, and expression of genetic information.
They are composed of nucleotide monomers, which consist of a sugar, a phosphate
group, and a nitrogenous base. The two main types of nucleic acids are:
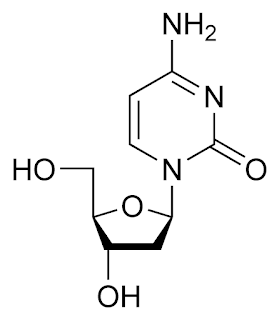
Deoxyribonucleic Acid (DNA)
1. It Stores genetic information in cells and has a double-helix structure.
2. It is considered to be a polynucleotides. One nucleotide unit consists of a deoxyribose sugar, four nitrogenous bases [adenine (A), thymine (T),
cytosine (C), and guanine (G)] and a phosphate group.
The nitrogenous bases are classified into two
types. They are purine (bicyclic ring named as adenine and guanine) and
pyrimidine (monocyclic rings named as cytosine, thymine, and uracil * Uracl
present in RNA)
A phosphate group is derived from phosphoric acid, with two of the hydrogen atoms from the phosphoric acid being replaced by bonds with the sugar molecules of the nucleotides.
Now that we have the idea of individual
component of a DNA, lets discuss further. When a nitrogenous base combines with
a ribose sugar it is called a nucleoside.
When a phosphate unit is added with the
Nucleoside it becomes the nucleotide. This nucleotide repeats itself to form a
complete chain of a DNA molecule.
1. Involved in protein synthesis and gene expression and is usually single-stranded.
2. It is composed of nucleotides with ribose
sugar and four nitrogenous bases [adenine (A), uracil (U), cytosine (C), and
guanine (G)].
A pairs with U, and C pairs with G.
Types of RNA: messenger RNA (mRNA),
transfer RNA (tRNA), and ribosomal RNA (rRNA).
Functions of Nucleic Acids:
DNA: Stores and transmits genetic
instructions.
RNA: Helps in protein synthesis and
gene regulation.
Carbohydrates: (*** Check whethr it is present in your University Syllabus, if not Skip this and scroll down to next Unit)
Carbohydrates are biological macromolecules made up of carbon (C), hydrogen (H), and oxygen (O), typically in a 1:2:1 ratio. They are one of the primary sources of energy for living organisms and also play structural and functional roles.
Types of Carbohydrates
Carbohydrates are classified into three main types based on their complexity:
Monosaccharides (Simple Sugars)
The basic building blocks of carbohydrates.
Examples:
Glucose (C₆H₁₂O₆) – Primary energy source.
Fructose – Found in fruits.
Galactose – Found in milk.
Disaccharides (Double Sugars)
Formed by the combination of two monosaccharides through a glycosidic bond.
Examples:
Sucrose (Glucose + Fructose) – Table sugar.
Lactose (Glucose + Galactose) – Milk sugar.
Maltose (Glucose + Glucose) – Found in germinating grains.
Polysaccharides (Complex Carbohydrates)
Long chains of monosaccharides used for energy storage or structural functions.
Examples:
Starch – Energy storage in plants.
Glycogen – Energy storage in animals (stored in liver and muscles).
Cellulose – Provides structural support in plant cell walls.
Chitin – Found in fungal cell walls and the exoskeleton of arthropods.
Functions of Carbohydrates
Energy Source: Glucose is the primary fuel for cellular respiration.
Energy Storage: Starch (plants) and glycogen (animals) store energy for later use.
Structural Role: Cellulose in plants and chitin in fungi and arthropods provide support.
Cell Communication: Some carbohydrates (glycoproteins, glycolipids) are involved in cell signaling and immune responses.
Configuration of Carbohydrates
Carbohydrates have specific structural configurations that determine their chemical properties and biological functions. These configurations are based on the arrangement of atoms and the orientation of functional groups in the molecule. The key aspects of carbohydrate configuration include isomerism, stereochemistry, and ring structures.
D annd L - Glucose:
In the open chain structure of glucose, when the -OH group at carbon number 5 or C5 (the chiral carbon with highest locant number) is towards right, that is refered to D-Glucose and when the same -OH group is towards left it is called L-glucose.
The d and l isomers of glucose are simply the optical isomers which rotate the plane of polarised light in opposite direction.
Anomers in Carbohydrates
Anomers are a specific type of stereoisomer found in cyclic carbohydrates. They differ in the configuration of the anomeric carbon (C-1 for aldoses, C-2 for ketoses) when a sugar forms a ring structure.
Below are the different repersentations of alpha-D-Glucopyranose as suggested by different chemists:
Mutarotation:
Mutarotation is the change in the optical rotation of a solution due to the interconversion between different anomers (α and β forms) of a sugar until equilibrium is reached. This phenomenon occurs in carbohydrates, especially in cyclic hemiacetals and hemiketals, such as glucose.
Example: Mutarotation of D-Glucose
Pure α-D-Glucose has an optical rotation of +112°. Pure β-D-Glucose has an optical rotation of +18.7°. When dissolved in water, the optical rotation changes until it stabilizes at +52.7°, representing the equilibrium mixture (about 36% α and 64% β).
Factors Affecting Mutarotation
1.Solvent: Water and polar solvents facilitate mutarotation.
2.pH: Acid or base catalysts speed up the interconversion.
3.Temperature: Higher temperatures increase the rate of mutarotation.
Importance of Mutarotation
1.Biological Systems: Enzymes recognize specific anomers, affecting digestion and metabolism.
2.Food Industry: Mutarotation influences sugar sweetness and crystallization.
3.Pharmaceuticals: Drug formulations involving sugars need to account for mutarotation effects.
Unit - II: Molecular interaction, thermodynamics, and kinetics of biological systems
Recombinant DNA Technology (Genetic Engineering)
Definition:
Recombinant DNA (rDNA)
technology involves joining DNA molecules from different sources and
inserting them into a host organism to produce new genetic
combinations or express specific proteins (like insulin, growth hormone,
etc.).
Key Steps in Recombinant
DNA Technology:
1.
Isolation of DNA:
o
Extract the DNA
containing the gene of interest from the donor organism.
2.
Cutting DNA with
Restriction Enzymes:
o
Use restriction
endonucleases to cut DNA at specific sequences (palindromic sites).
o
Produces sticky or
blunt ends.
3.
Insertion into Vector:
o
The DNA fragment is
inserted into a vector (like plasmid, bacteriophage, or cosmid).
o
Ligase enzyme joins the DNA fragments.
4.
Introduction into Host
Cell:
o
Transfer the recombinant
vector into a host organism (commonly E. coli) via transformation,
transduction, or electroporation.
5.
Selection and Screening:
o
Use antibiotic
resistance genes or reporter genes to identify recombinant organisms.
o
Blue-white screening
(lacZ gene) is commonly used.
6.
Expression of the Gene:
o
The host transcribes and
translates the inserted gene, producing the desired protein.
7.
Protein Harvesting:
o
The expressed recombinant
protein is extracted and purified.
Applications of rDNA
Technology:
·
Production of therapeutic
proteins (e.g., insulin, HGH)
·
Gene therapy
·
GMOs in agriculture
·
Vaccine development
(e.g., Hepatitis B)
·
Study of gene
functions and regulation
2. Protein Purification
Definition:
Protein purification is the process of isolating a
specific protein from a complex mixture for research or therapeutic use.
Steps in Protein
Purification:
1.
Cell Lysis:
o
Break open cells using
mechanical, enzymatic, or chemical methods.
2. Crude Extraction:
o
Separate soluble proteins
from cell debris by centrifugation.
3. Purification Techniques (Based on
physicochemical properties):
a. Precipitation:
o
Use salts (like ammonium
sulfate) or pH to precipitate proteins.
b. Chromatography:
o
Ion-exchange
chromatography (based on charge)
o
Gel filtration
chromatography (based on size)
o
Affinity chromatography (based on specific binding, e.g., His-tag/Ni column)
c. Electrophoresis:
o
SDS-PAGE for checking
purity and estimating molecular weight.
d. Ultracentrifugation:
o
Based on size and density
for high-resolution separation.




.png)









I was astonished to that Sharukh Khan has a relation with chemistry. Wah.
ReplyDeleteWaao , nice to see our celebrities loved chemistry
ReplyDeleteSir send interesting topics like this
ReplyDelete